About
FerrDb is the world's first manually curated database for ferroptosis regulators and ferroptosis-disease associations from published journal articles.
This major update, FerrDb V3, expands FerrDb to a database for 22 regulated cell death (RCD) modalities. FerrDb is well-maintained and regularly updated to support long-term service. The data in FerrDb is free to use for academic purposes.
RCD is a meticulously controlled process that orchestrates the elimination of cells through specific signaling pathways, playing a vital role in maintaining organismal homeostasis, development, and immune defense. RCD is essential for removing damaged, infected, or potentially harmful cells, thus preventing diseases such as cancer and autoimmune disorders. However, dysregulation of RCD pathways can result in pathological conditions, highlighting the importance of understanding these processes for therapeutic interventions.
FerrDb dedicates to RCD regulators and RCD-disease associations. There are two secondary categories of RCD regulators: (1) genes and (2) substances. Gene regulators include driver, suppressor, marker, and unclassified gene. Substances cover a wide range of chemical entities, including pure substances (e.g., iron, erastin) and mixtures (e.g., herbal extracts). Substance regulators include inducers and inhibitors. On the basis of this philosophy, FerrDb consists of seven individual curated datasets.
We are in an era of knowledge explosion, making it crucial to find efficient ways to organize, utilize, and share knowledge. This update aims to meet the demands of rapidly advancing RCD research.
RCD coverage
| RCD | Publication date range | Curation strategy | Article count |
|---|---|---|---|
| ADCD | Up to 2025-05-31 | Manual curation | 184 |
| Autosis | Up to 2025-05-31 | Manual curation | 68 |
| Alkaliptosis | Up to 2025-05-31 | Manual curation | 54 |
| Apoptosis | Not applicable | From Gene Ontology | NA |
| Intrinsic apoptosis | Not applicable | From Gene Ontology | NA |
| Mitotic death | Up to 2025-05-31 | Manual curation | 131 |
| Extrinsic apoptosis | Not applicable | From Gene Ontology | NA |
| Entotic cell death | Up to 2025-05-31 | Manual curation | 214 |
| Ferroptosis | Up to 2024-12-31 | Manual curation | 18371 |
| Immunogenic cell death | Up to 2025-05-31 | Manual curation | 4507 |
| LDCD | Up to 2025-05-31 | Manual curation | 88 |
| NETotic cell death | Up to 2025-05-31 | Manual curation | 25 |
| Regulated necrosis | Up to 2025-05-31 | Manual curation | 436 |
| Necroptosis | Not applicable | From Gene Ontology | NA |
| MPT-driven necrosis | Up to 2025-05-31 | Manual curation | 95 |
| Oxeiptosis | Up to 2025-05-31 | Manual curation | 64 |
| Parthanatos | Up to 2025-05-31 | Manual curation | 408 |
| Pyroptosis | Not applicable | From Gene Ontology | NA |
| PANoptosis | Up to 2024-06-17 | Manual curation | 291 |
| Cuproptosis | Up to 2023-12-31 | Manual curation | 831 |
| Disulfidptosis | Up to 2025-05-31 | Manual curation | 461 |
| Spectosis | Up to 2025-05-31 | Manual curation | 1 |
Data structure
| Dataset | Primary category | Secondary category | Tertiary category | Description |
|---|---|---|---|---|
| Driver | Regulator | Gene | Driver | A gene that promotes RCD. |
| Suppressor | Regulator | Gene | Suppressor | A gene that prevents RCD. |
| Marker | Regulator | Gene | Marker | A gene that indicates RCD occurrence. |
| Unclassified gene | Regulator | Gene | Unclassified | A gene that is associated with RCD but its regulatory role is unclear. |
| Inducer | Regulator | Substance | Inducer | A substance entity which enables to cause RCD. |
| Inhibitor | Regulator | Substance | Inhibitor | A substance entity which enables to restrict RCD. |
| RCD-disease association | Association | Disease association | Disease | RCD has two opposite effects on disease: aggravating an illness or alleviating an unpleasant situation. |
Confidence level
| Confidence | Explanation |
|---|---|
| Validated | The most reliable. It requires convincing evidence from strict tests such as pharmacological or genetic inhibition or activation test. |
| Screened | No strict validation. Only high-throughput data is available. |
| Predicted | No strict validation. In the source article, the authors drew their conclusions based on computational results or their knowledge. |
| Deduced | No strict validation. We, the curators, inferred the gene's function based on our understanding of the source article. |
| GO | An entry from the GO database. |
Browse data
Click Browse in the navigation bar to open the browse page.
Data entries are shown on the page in a similar format. As shown in the following graph, each row is a regulator and attributes are shown in columns.
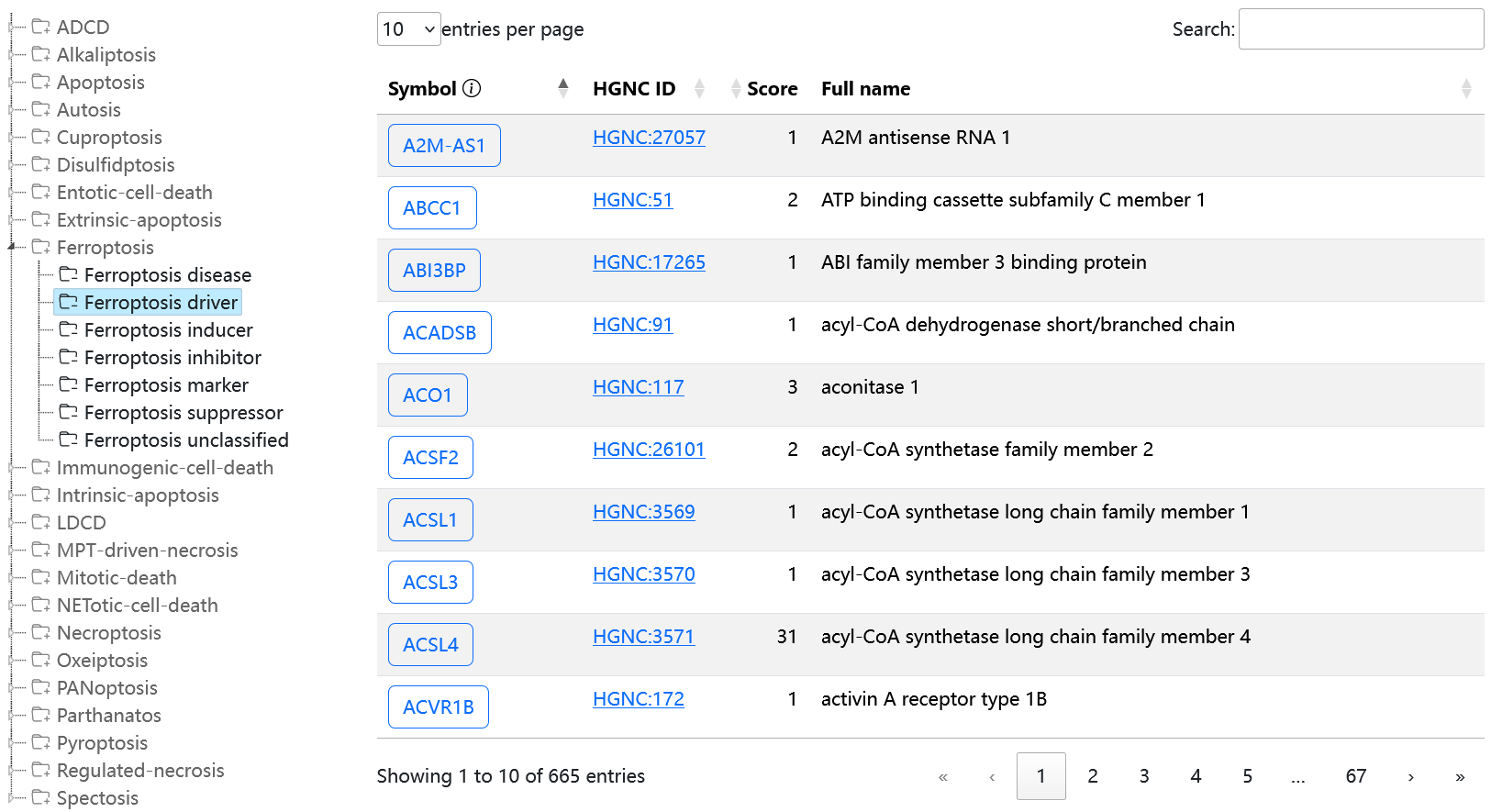
Click the symbol of a gene in the first column to open the gene detail page.
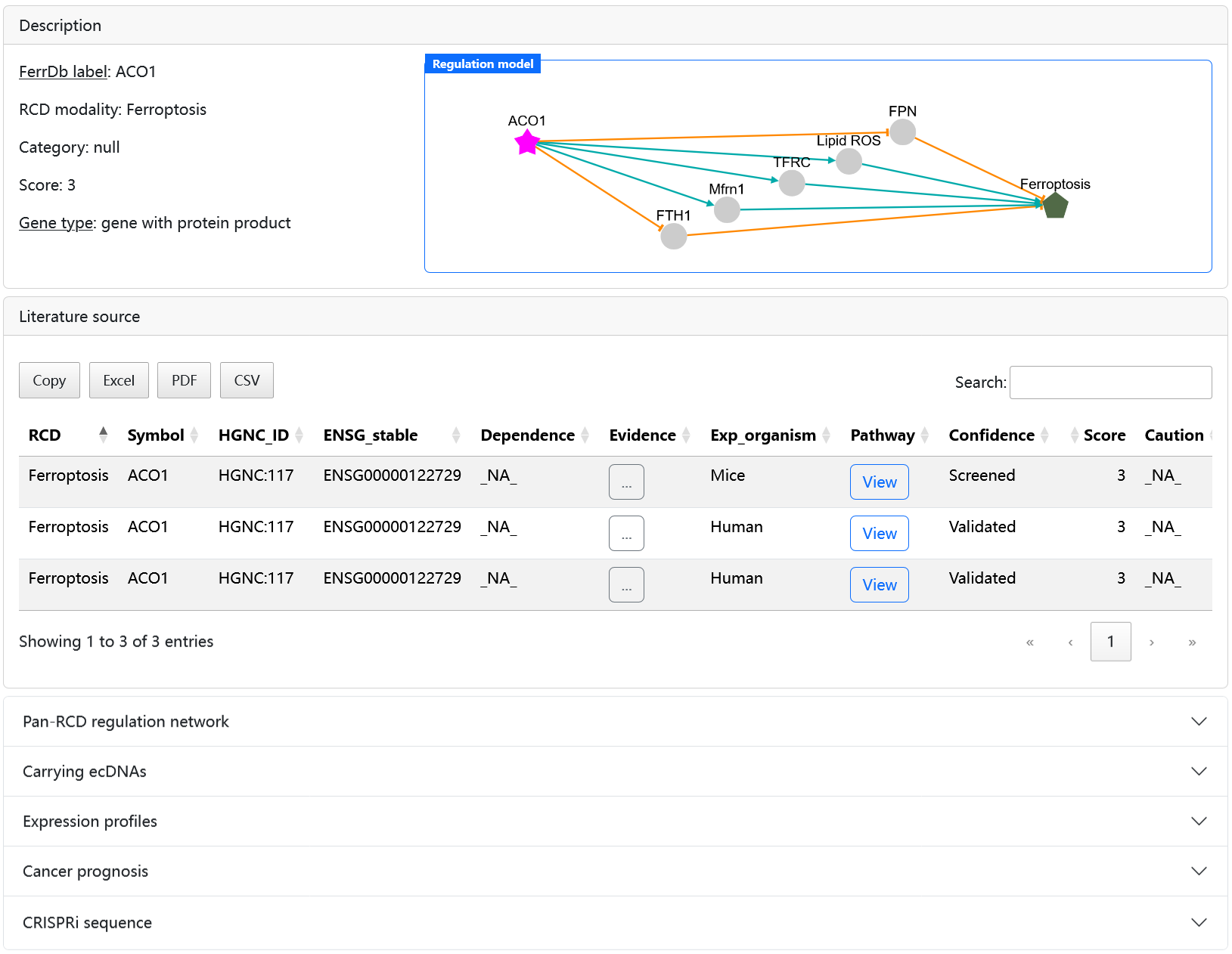
The Last_update column shows when the item was last updated. Please be notified that for items already in FerrDb V1, the update date is set to 2020/12/31. As of FerrDb V2, the date is set to the date of its inclusion or modification.
Diagram legend
Chatbot
Disclaimer: The outputs of ChatRCD are generated automatically and are intended for informational purposes. They are not guaranteed for accuracy and should be independently verified before use in any case.
ChatRCD is a specialized chatbot designed to provide intuitive, question-and-answer-based interaction with FerrDb. It is built on a Retrieval-Augmented Generation (RAG) framework, leveraging the powerful Qwen-Turbo (Qwen3) foundation model. To ensure domain-specific accuracy, our manually curated RCD data has been converted into a vectorized knowledge base for the RAG retriever. This architecture enables ChatRCD to retrieve highly relevant information and generate contextually accurate answers to complex queries about RCD.
Example prompts
- List five ferroptosis suppressor genes.
- Can dulaglutide inhibit LDCD?
- Sorafenib enables to induce both ferroptosis and cuproptosis. Right?
- Dose autosis has an effect on Sjögren's disease?
- Which gene can suppress apoptosis: MIR27A, PI3KCA, PAX2, CDK6?
- What is the evidence supporting IL1R1 as a suppressor of regulated necrosis?
- What are the multiple roles of HMOX1 in regulating ferroptosis?
- What types of RCD can be regulated by ATG5?
Download data
Please go to the Download page to get the data. The data are released under a Creative Commons Attribution-NonCommercial 4.0 International (CC BY-NC 4.0) License.
Submit data
We appreciate your contribution. Please go to the Submit page and follow the guidlines on that page to submit data to FerrDb.
Use utilities
Acceptable ID types
Analytical utilities accept the ID types listed in order of priority in the following. They will take care of the ID conversions for non-symbol IDs.
- HGNC symbol [Recommended] (eg., A1BG, A2M, A2ML1-AS1, ...)
- HGNC ID (eg., HGNC:5, HGNC:7, HGNC:41022, ...)
- Ensembl gene stable ID (eg., ENSG00000210049, ENSG00000211459, ENSG00000210077, ...)
- Ensembl gene ID version (eg., ENSG00000210049.1, ENSG00000211459.2, ENSG00000210077.12, ...)
- NCBI gene ID (eg., 29974, 2, 144568, ...)
- RefSeq gene ID (eg., NG_029916.1, NG_011717.2, NG_042857.2, ...)
- UniProtKB accession (eg., A0A023HJ61, A0A023HN28, A0A023I7F4, ...)
- UniProtKB ID (eg., A0A023HJ61_HUMAN, A0A023HN28_HUMAN, A0A023I7F4_HUMAN, ...)
- GI (eg., 498922526, 511106493, 540070669, ...)
vennRCD
This tool can be used for Venn analysis.
Input
- A gene list
Parameters
- RCD
- Category
Output
- Venn plots and overlapping subsets.
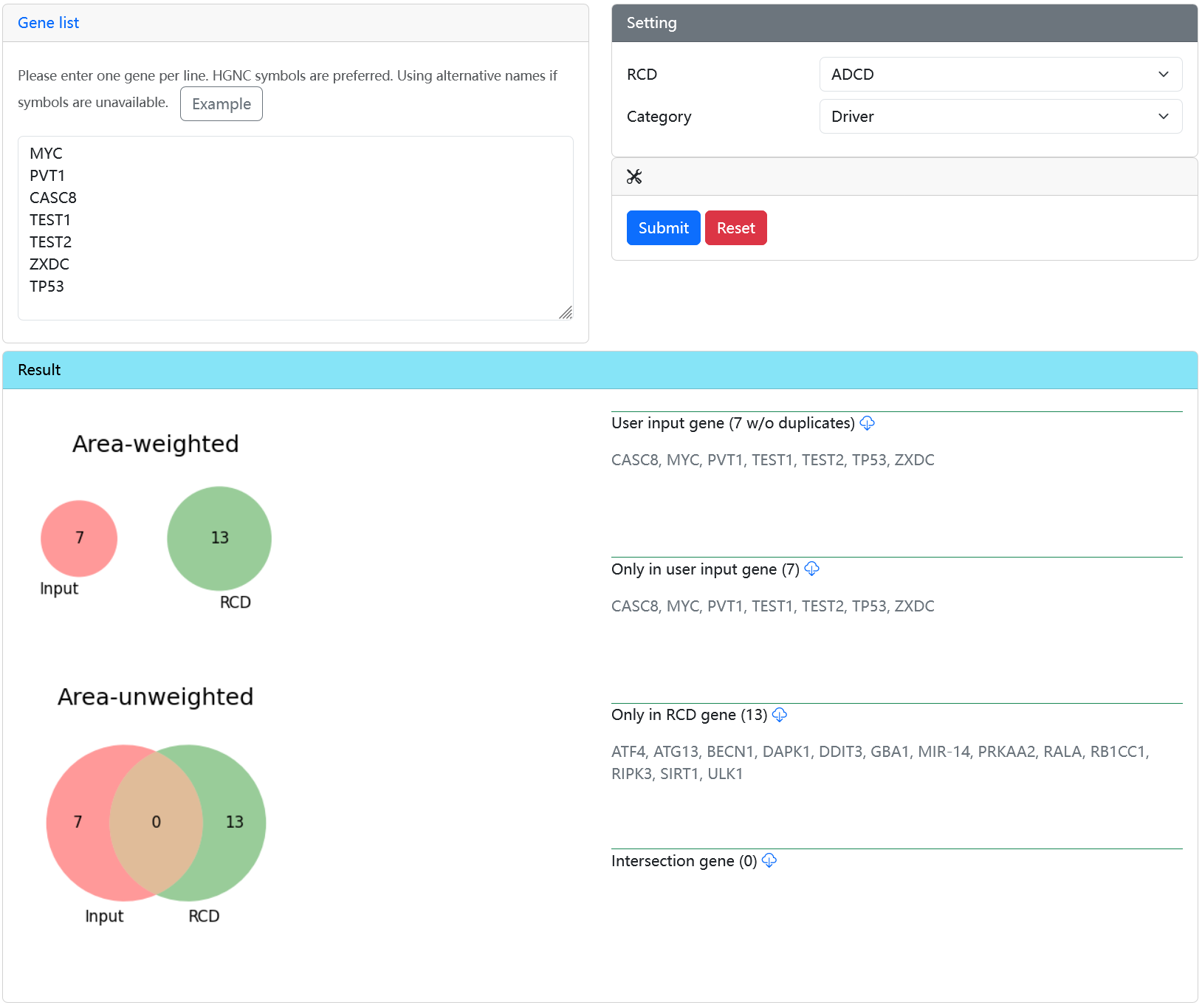
enrichRCD
This enrichment analysis tool enables users to know whether a gene list of their interest is statistically enriched in any RCD gene set annotated in FerrDb. The analysis is performed using GSEApy, with both Enrichr's over-representation and Subramanian et al.'s GSEA methods available.
Input
- A single-column gene list
- A double-column gene list
- An expression table and a condition file
Output
- Enrichment plot and result table.
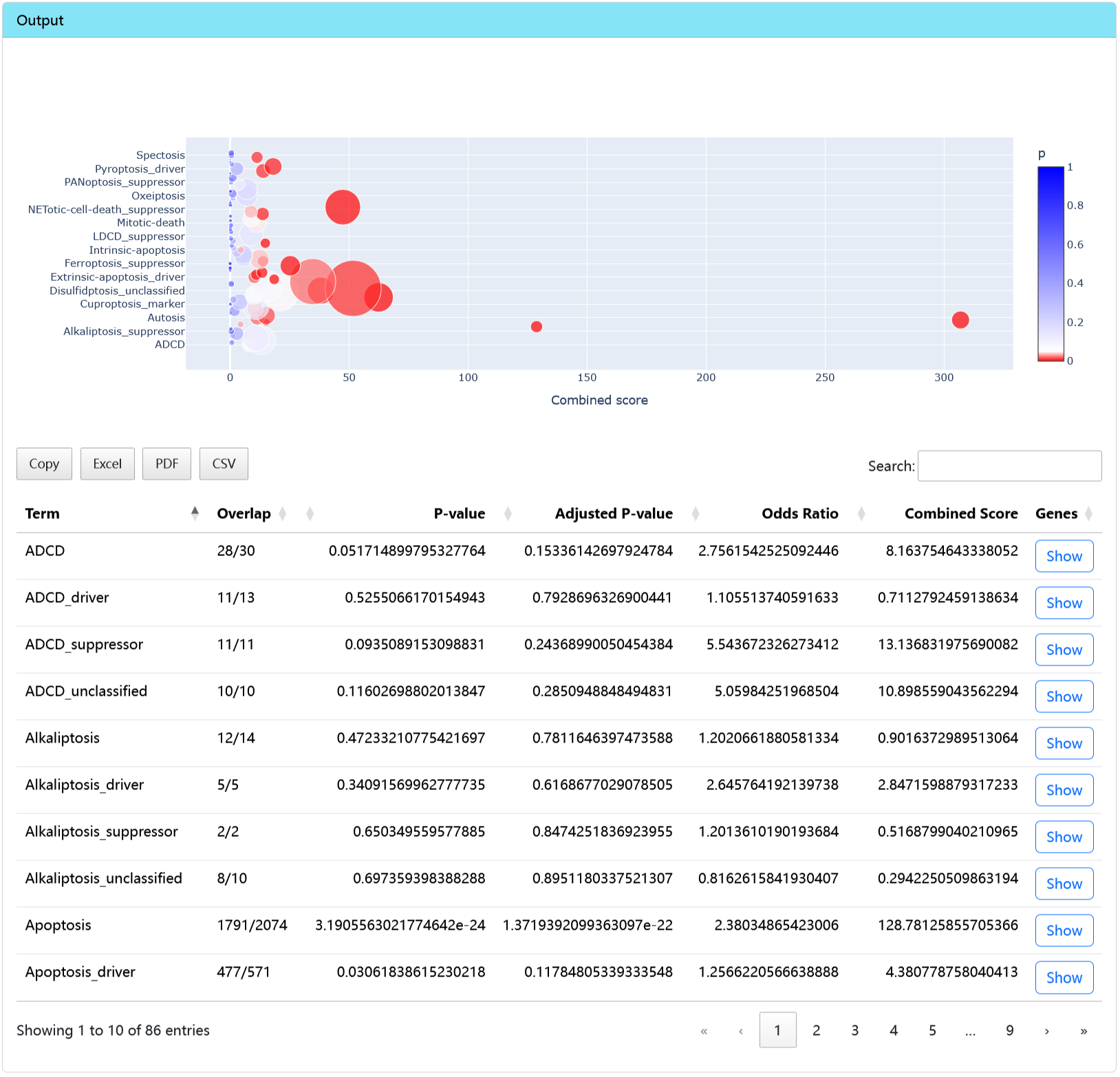
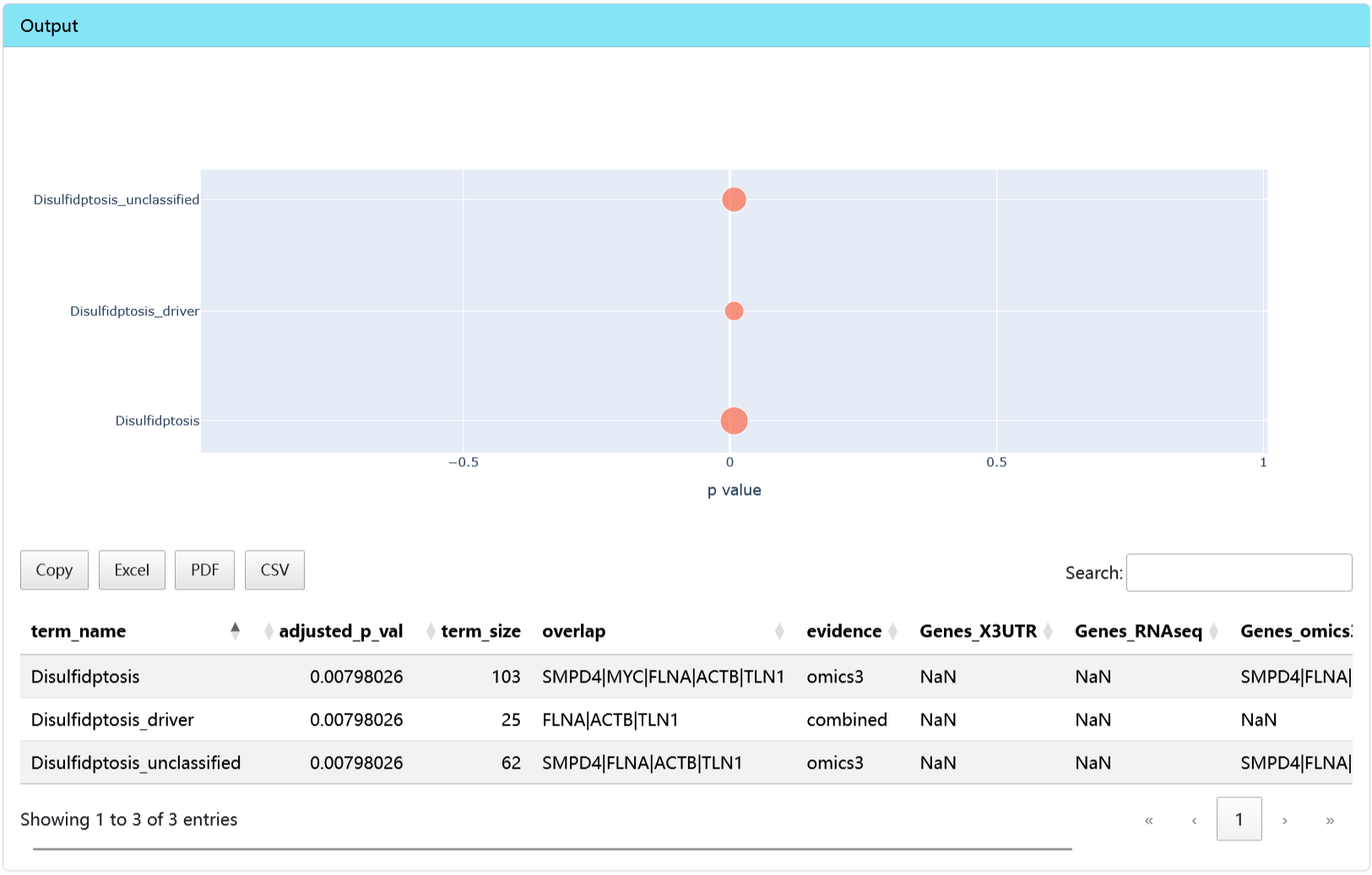
omicsRCD
This tool can be used for enrichment analysis based on multi-omics datasets.
Input
- A table of concatenated multi-omics datasets.
Output
- Result plot and table.
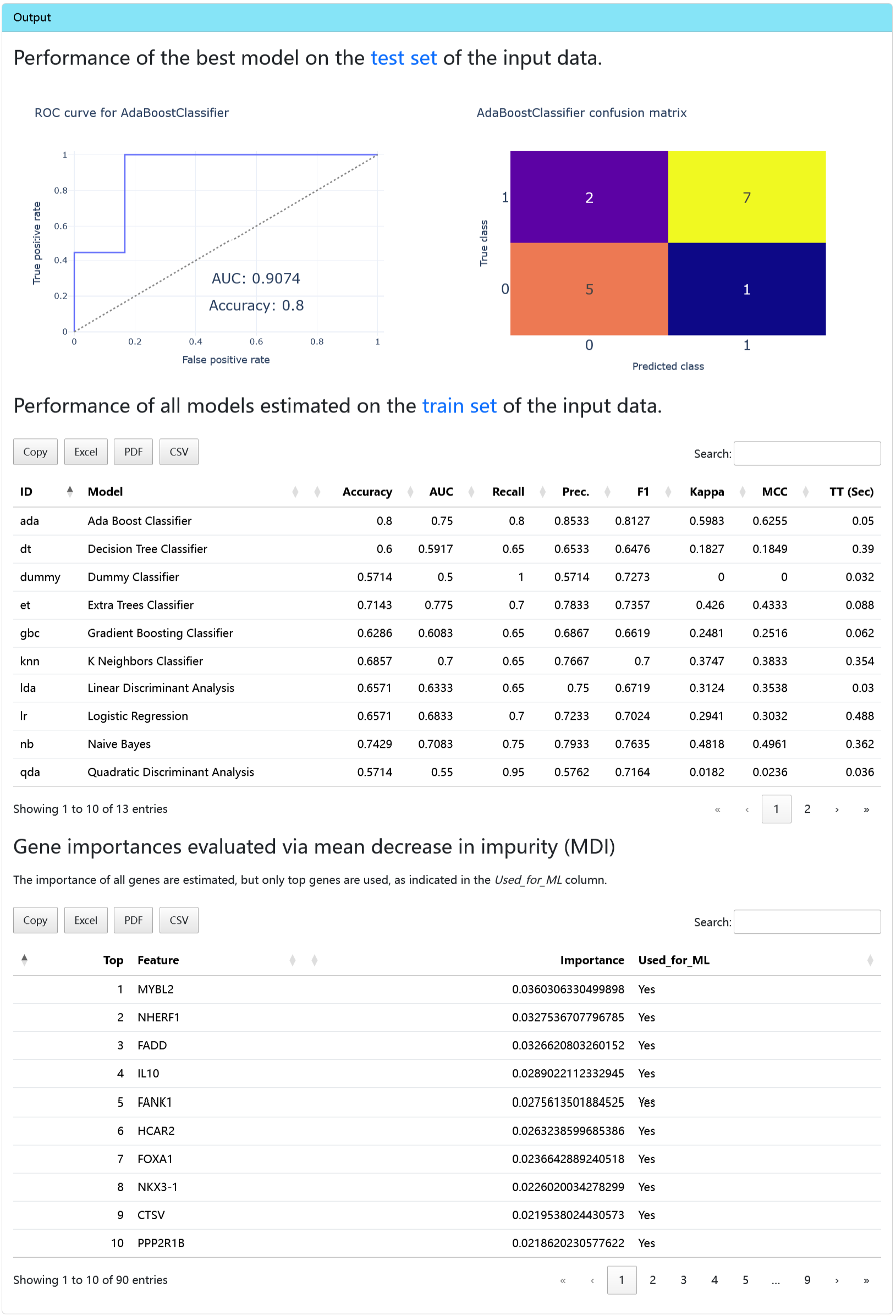
classRCD
This tool can be used for machine learning-based classification tasks.
Input
- A data file and a condition file
Parameters
- RCD
- Category
- Number of top important genes
Output
- Performance plots
- Model metrics
- Gene importance table
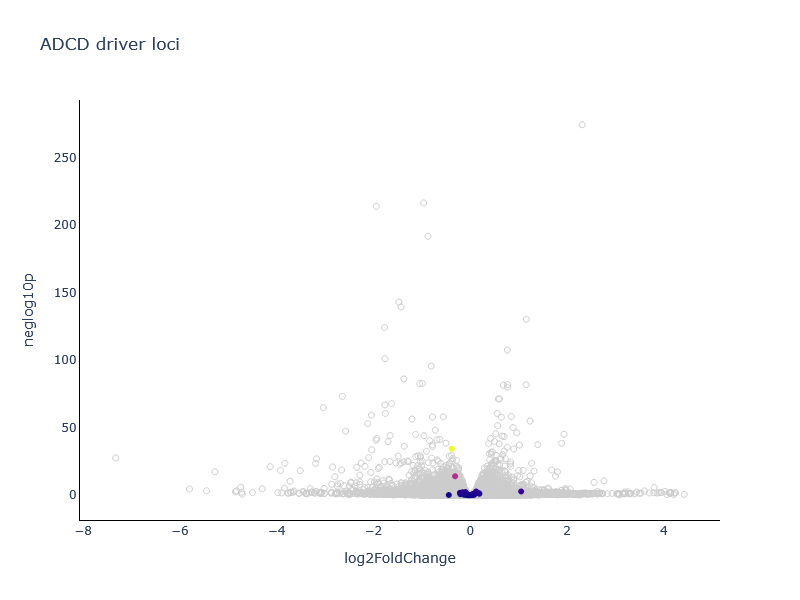
locateRCD
This tool can be used to reveal significant RCD genes on overplotted diagrams.
Input
- A file with two or three columns
Parameters
- RCD
- Category
Output
- A plot with RCD genes highlighted
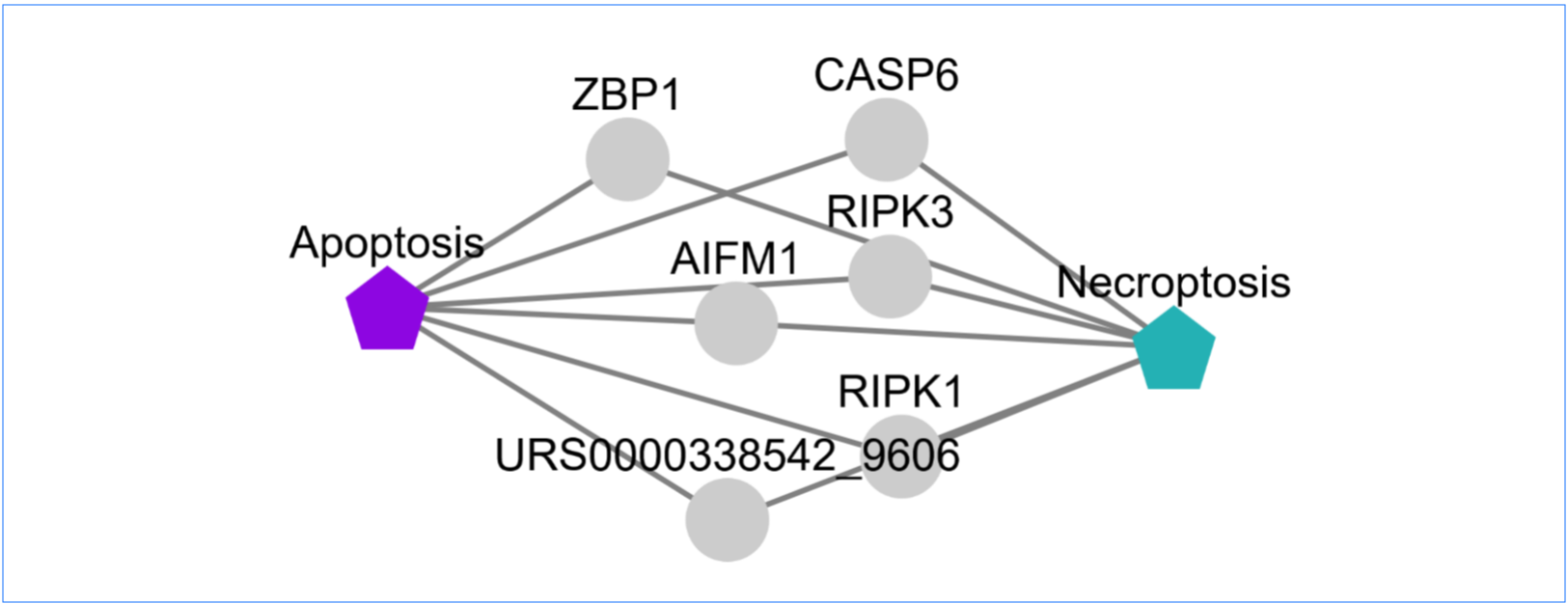
crossRCD
This tool can be used for network visualization of the crosstalk between two RCD modalities.
Parameters
- RCD 1
- RCD 2
- Category
Output
- Interaction network
Feedback
We'd love to hear from you! If you have any comments, please email the webmaster at admin@zhounan.org.
Working group
| Name | Task/Role | Affiliation |
|---|---|---|
| Nan Zhou | Article collection | The Affiliated Brain Hospital, Guangzhou Medical University |
| Curation and annotation | ||
| Coding | ||
| Coordinator | ||
| Yuping Ning | Quality control | The Affiliated Brain Hospital, Guangzhou Medical University |
| Qiqi Luo | Curation and annotation | College of Life Sciences, Sichuan University |
| Tong Yin | Curation and annotation | Sun Yat-Sen Memorial Hospital, Sun Yat-Sen University |
| Jinku Bao | Quality control | College of Life Sciences, Sichuan University |
| Xiaoqing Yuan | Project management | Sun Yat-Sen Memorial Hospital, Sun Yat-Sen University |
| Quality control | ||
| Li Peng | Data management | Sun Yat-Sen Memorial Hospital, Sun Yat-Sen University |
| Curation and annotation |
Image credit
Cell illustrations on the home page are created with BioGDP.com.